Systems of Cell Reproduction
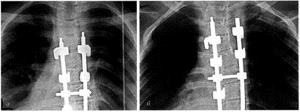
Unicellular organisms use cell division primarily to reproduce themselves, whereas in multicellular organisms cell division also plays important roles in growth and in the repair of tissues (Fig. 2.5). In order for any cell to divide, four events must occur: There must be a reproductive signal. This signal, which may come either from inside or outside the cell, initiates the cellular reproductive events.
Replicationof DNA (the genetic material) and other vital cell components must occur so that each of the two new cells will be identical and have complete cell functions.
The cell must distribute the replicated DNA to each of the two new cells. This process is called segregation.
New material must be added to the cell membrane (and the cell wall, in organisms that have one) in order to separate the two new cells in a process called cytokinesis.
These four events occur in somewhat different ways in prokaryotes and eukaryotes.
Prokaryotes divide by fission
In prokaryotes, cell division results in the reproduction of the entire single-celled organism. The cell grows in size, replicates its DNA, and then essentially divides into two new cells, a process called fission.
Reproductive signals
The reproductive rates of many prokaryotes respond to conditions in the environment. The bacterium Escherichia coli, a species that is commonly used in genetic studies, is a “cell division machine” that essentially divides continuously. Typically, cell division takes 40 minutes at 37°C. But if there are abundant sources of carbohydrates and salts available, the division cycle speeds up so that cells may divide in 20 minutes. Another bacterium, Bacillus subtilis, stops dividing when food supplies are low, then resumes dividing when conditions improve. These observations suggest that external factors, such as materials in the environment, control the initiation of cell division in prokaryotes.
Replication of DNA
A chromosome, is a DNA molecule containing genetic information. When a cell divides, all of its chromosomes must be replicated, and each of the two resulting copies must find its way into one of the two new cells.
Most prokaryotes have only one chromosome, a single long DNA molecule with proteins bound to it. In the bacterium E. coli, the DNA is a continuous molecule often referred to as a circular chromosome. If the bacterial DNA were actually arranged in a circle, it would be about 1.6 million nm (1.6 mm) in circumference. Thus the bacterial DNA, fully extended, would form a circle over 100 times larger than the cell! To fit it into the cell, the DNA must be packaged. The DNA molecule accomplishes some packaging by folding in on itself, and positively charged (basic) proteins bound to negatively charged (acidic) DNA contribute to this folding. Circular chromosomes appear to be characteristic of all prokaryotes, as well as some viruses, and are also found in the chloroplasts and mitochondria of eukaryotic cells. Functionally, the prokaryotic chromosome has two regions that are important for cell reproduction:
The site where replication of the circle starts: the origin of replication, designated ori (entation)
The site where replication ends: the terminus of replication, ter
The process of chromosome replication occurs as the DNA is threaded through a “replication complex” of proteins at the center of the cell. These proteins include the enzyme DNA polymerase. During the process of prokaryotic DNA replication, the cell grows and provides a mechanism for the ordered distribution of the DNA into the newly formed daughter cells.
Segregation of DNA
DNA replication actively drives the segregation of the replicated DNA molecules to the two new cells. The first region to be replicated is ori, which is attached to the plasma membrane. The two resulting ori regions separate as the new chromosome forms and new plasma membrane forms between them as the cell grows longer. By the end of replication, there are two chromosomes, one at either end of the lengthened bacterial cell.
Cytokinesis
Cell separation, or cytokinesis, begins 20 minutes after chromosome replication is finished. The first event of cytokinesis is a pinching in of the plasma membrane to form a ring similar to a purse string. Fibers composed of a protein similar to eukaryotic tubulin are major components of this ring. As the membrane pinches in, new cell wall materials are synthesized which finally separate the two cells.
Eukaryotic cells divide by mitosis or meiosis
Complex eukaryotes, such as humans and flowering plants, originate from a single cell, the fertilized egg. This cell derives from the union of two sex cells, called gametes, from the organism’s parents—that is, a sperm and egg—and so contains genetic material from both of these parental cells. This means that the fertilized egg contains one set of chromosomes from the male parent and one set from the female parent.
The formation of a multicellular organism from a fertilized egg is called development. It involves both cell reproduction and cell specialization. For example, an adult human has several trillion cells, all ultimately deriving from the fertilized egg, and many of them have specialized roles. We will discuss how cells specialize later in this book, in Part Three. For now, we will focus on cell reproduction. Cell reproduction in eukaryotes, like that in prokaryotes, involves reproductive signals, DNA replication, segregation, and cytokinesis. But, as you might expect, events in eukaryotes are somewhat more complex. First, unlike prokaryotes, eukaryotic cells do not constantly divide whenever environmental conditions are adequate. In fact, eukaryotic cells that are part of a multicellular organism and have become specialized seldom divide. So the signals for cell division are related not to the environment of a single cell but to the needs of the entire organism. Second, instead of a single chromosome, eukaryotes usually have many (humans have 46), so the processes of replication and segregation, while basically the same as in prokaryotes, are more intricate. Third, eukaryotic cells have a distinct nucleus, which has to be replicated and then divided into two new nuclei. Thus, in eukaryotes, cytokinesis is distinct from division of the genetic material. Finally, cytokinesis is different in plant cells (which have a cell wall) than in animal cells (which do not). The key difference between prokaryotic and eukaryotic cell reproduction is that in the eukaryotes, newly replicated chromosomes remain associated with each other as sister chromatids and a new mechanism, mitosis, is used to segregate them into the two new nuclei.
The reproduction of a eukaryotic cell typically consists of three steps:
The replication of DNA within the nucleus
The packaging and segregation of the replicated DNA into two new nuclei (nuclear division)
The division of the cytoplasm (cytokinesis)
A second mechanism of nuclear division, meiosis, occurs in germ cells that produce gametes that contribute to the reproduction of a new organism. While the two products of mitosis are genetically identical to the cell that produced them— they both have the same DNA—the products of meiosis are not. As we will see later in the chapter, meiosis generates diversity by shuffling the genetic material, resulting in new gene combinations. It plays a key role in sexual life cycles.
Interphase and the Control of Cell Division
A cell lives and functions until it divides or dies. Or, if it is a gamete, it lives until it fuses with another gamete. Some types of cells, such as red blood cells, muscle cells, and nerve cells, lose the capacity to divide as they mature. Other cell types, such as cortical cells in plant stems, divide only rarely. Some cells, like the cells in a developing embryo, are specialized for rapid division. Between divisions—that is, for most of its life—a eukaryotic cell is in a condition called interphase. For most types of cells, we may speak of a cell cyclethat has two phases: mitosis and interphase. In this section, we will describe the cell cycle events that occur during interphase, especially the “decision” to enter mitosis.
A given cell lives for one turn of the cell cycle and then becomes two cells. The cell cycle, when repeated again and again, is a constant source of new cells. However, even in tissues engaged in rapid growth, cells spend most of their time in interphase. Examination of any collection of dividing cells, such as the tip of a root or a slice of liver, will reveal that most of the cells are in interphase most of the time; only a small percentage of the cells will be in mitosis at any given moment. Interphase consists of three subphases identified as G1, S, and G2. The cell’s DNA replicates during the S phase(the S stands for synthesis). The period between the end of mitosis and the onset of the S phase is called G1 or Gap 1. Another gap phase—G2—separates the end of the S phase and the beginning of mitosis, when nuclear and cytoplasmic division take place and two new cells are formed. Mitosis and cytokinesis are referred to as the M phaseof the cell cycle (Figure ). The process of DNA replication is completed by the end of S phase. Where there was formerly one chromosome, there are now two, joined together and awaiting segregation into two new cells by mitosis or meiosis. Although one key event—DNA replication—dominates and defines the S phase, important cell cycle processes take place in the gap phases as well. G1 is quite variable in length in different cell types. Some rapidly dividing embryonic cells dispense with it entirely, while other cells may remain in G1 for weeks or even years. In many cases, these cells enter a resting phase called G0. Special internal and external signals are needed to prompt a cell to leave G0 and re-enter the cell cycle at G1. The biochemical hallmark of a G1 cell is that it is preparing for the S phase, so at this stage each chromosome is a single, unreplicated structure. It is at the G1-to-S transition that the commitment to enter another cell cycle is made. During G2, the cell makes preparations for mitosis—for example, by synthesizing components of the microtubules that will move the chromosomes to opposite ends of the dividing cell. Because the chromosomes were replicated during the S phase, each chromosome now consists of two identical sister chromatids.
Cyclins and other proteins signal events in the cell cycle
How are appropriate decisions to enter the S or M phases made? These transitions—from G1 to S and from G2 to M— depend on the activation of a type of protein called cyclin-dependent kinase, or Cdk. Remember that a kinase is an enzyme that catalyzes the transfer of a phosphate group from ATP to another molecule; this phosphate transfer is called phosphorylation:
Kinase protein + ATP → protein — P + ADP
What does phosphorylation do to a protein? The proteins have both hydrophilic regions (which tend to interact with water on the outside of the macromolecule) and hydrophobic regions (which tend to interact with one another on the inside of the macromolecule). These regions are important in giving a protein its three-dimensional shape. Phosphate groups are charged, so an amino acid with such a group tends to be on the outside of the protein. In this way, phosphorylation changes the shape and function of a protein. By catalyzing the phosphorylation of certain target proteins, Cdk’s play important roles in initiating the steps of the cell cycle. The discovery that Cdk’s induce cell division is a beautiful example of research on different organisms and different cell types converging on a single mechanism:
One group of scientists was studying immature sea urchin eggs, trying to find out how they are stimulated to divide and form mature eggs. A protein called maturation promoting factor was purified from maturing eggs, which by itself prodded immature eggs into division.
Other scientists studying the cell cycle in yeast, a singlecelled eukaryote, found a strain that was stalled at the G1–S boundary because it lacked a Cdk. This yeast Cdk was discovered to be very similar to the sea urchin’s maturation promoting factor.
Similar Cdk’s were soon found to control the G1-to-S transition in many other organisms, including humans. But Cdk’s are not active by themselves. Rather, they must bind to a second type of protein, called cyclin. This binding— an example of allosteric regulation—activates the Cdk by altering its shape and exposing its active site (see Figure ). It is the cyclin-Cdk complex that acts as a protein kinase and triggers the transition from G1 to S phase. Then cyclin breaks down, and the Cdk becomes inactive. Several different cyclin-Cdk combinations act at various stages of the mammalian cell cycle:
Cyclin D-Cdk4 acts during the middle of G1. This is the restriction point (R), a key decision point beyond which the rest of the cell cycle is normally inevitable.
Cyclin E-Cdk2 also acts in the middle of G1.
Cyclin A-Cdk2 acts during S, and also stimulates DNA replication.
Cyclin B-Cdk1 acts at the G2–M boundary, initiating the transition to mitosis.
The key to progress past the restriction point is a protein called RB (retinoblastoma protein, named for a childhood cancer in which it was first discovered). RB normally inhibits the cell cycle. But when RB is phosphorylated by a protein kinase, it becomes inactive and no longer blocks the restriction point and the cell progresses past G1 into S phase (Note the double negative here—a cell function happens because an inhibitor is inhibited! This phenomenon is rather common in the control of cellular metabolism.) The enzymes that catalyze RB phosphorylation are Cdk4 and Cdk2. So what is needed for a cell to pass the restriction point is the synthesis of cyclins D and E which activate Cdk 4 and 2, which phosphorylate RB which becomes inactivated. The cyclin-Cdk complexes act as checkpoints, points at which a cell cycle’s progress can be monitored to determine whether the next step can be taken. For example, if DNA is damaged by radiation during G1, a protein called p21 is made. (The p stands for “protein” and the 21 stands for its molecular weight—about 21,000 daltons.) The p21 protein then binds to the two G1 Cdk’s, preventing their activation by cyclins. So the cell cycle stops while repairs are made to DNA. The p21 protein itself breaks down after the DNA is repaired, allowing cyclins to bind to the Cdk’s and the cell cycle to proceed. Because cancer results from inappropriate cell division, it is not surprising that these cyclin-Cdk controls are disrupted in cancer cells. For example, some fast-growing breast cancers have too much cyclin D, which overstimulates Cdk4 and thus cell division. A major protein in normal cells that prevents them from dividing is p53, which leads to the synthesis of p21 and therefore inhibition of Cdk’s. More than half of all human cancers contain defective p53 resulting in the absence of cell cycle controls.
Growth factors can stimulate cells to divide
Cyclin-Cdk complexes provide an internal control on progress through the cell cycle. But there are tissues in the body in which cells no longer go through the cell cycle or go through it slowly and divide infrequently. If such cells are to divide, they must be stimulated by external signals (chemical messengers) called growth factors. For example, when you cut yourself and bleed, specialized cell fragments called platelets gather at the wound and help to initiate blood clotting. The platelets also produce and release a protein, called platelet-derived growth factor, that diffuses to the adjacent cells in the skin and stimulates them to divide and heal the wound. Other growth factors include interleukins, which are made by one type of white blood cell and promote cell division in other cells that are essential for the body’s immune system defenses. Erythropoietin, made by the kidney, stimulates the division of bone marrow cells and the production of red blood cells. In addition, many hormones promote division in specific cell types. We will describe the physiological roles of growth factors in later chapters, but all of them act in a similar way. They bind to their target cells via specialized receptor proteins on the target cell surface. This specific binding triggers events within the target cell that initiate the cell cycle. Cancer cells often divide inappropriately because they make their own growth factors or because they no longer require growth factors to start cycling.
Eukaryotic Chromosomes
Most human cells other than gametes contain two full sets of genetic information, one from the mother and the other from the father. As in prokaryotes, this genetic information consists of DNA molecules. However, unlike prokaryotes, humans and other eukaryotes have more than one chromosome and during interphase those chromosomes reside within a membrane-enclosed organelle, the nucleus. The basic unit of the eukaryotic chromosome is a gigantic, linear, double-stranded molecule of DNA complexed with many proteins to form a dense material called chromatin. Before the S phase, each chromosome contains only one such double-stranded DNA molecule. However, after the DNA molecule replicates during the S phase, the two resulting DNA molecules, now called chromatids, are held together along most of their length by a protein called cohesin. They stay this way until mitosis, when most of the cohesin is removed, except in a region called the centromereat which the chromatids are still held together (see Figure). A second group of proteins called condensinscoats the DNA molecules at this time and makes them more compact. The DNA in a typical human cell has a total length of 2 meters. Yet the nucleus is only 5 m (0.000005 meters) in diameter. So, although the DNA in an interphase nucleus is “unwound,” it is still impressively packed! This packing is achieved largely by proteins associated closely with the chromosomal DNA. Chromosomes contain large quantities of proteins called histones. There are five classes of histones. All of them have a positive charge at cellular pH levels because of their high content of the basic amino acids lysine and arginine. These positive charges electrostatically attract the negative phosphate groups on DNA. These interactions, as well as interactions among the histones themselves, form beadlike units called nucleosomes. Each nucleosome contains the following components:
Eight histone molecules, two each of four of the histone classes, united to form a core or spool. 146 base pairs of DNA, 1.65 turns of it wound around the histone core.
Histone H1 (the remaining histone class) on the outside of the DNA, which may clamp it to the histone core.
During interphase, a chromosome is made up of a single DNA molecule running around vast numbers of nucleosomes like beads on a string. Between the nucleosomes stretches a variable amount of non-nucleosomal “linker” DNA. Since this DNA is exposed to the nuclear environment, it is accessible to proteins involved in its duplication and the regulation of its expression. During both mitosis and meiosis, the chromatin becomes ever more coiled and condensed as its nucleosomes pack together and coil, with further folding of the chromatin continuing up to the time at which the chromosomes begin to move apart.
Mitosis: Distributing Exact Copies of Genetic Information
In mitosis, a single nucleus gives rise to two nuclei that are genetically identical to each other and to the parent nucleus.
This process ensures the accurate distribution of the eukaryotic cell’s multiple chromosomes to the daughter nuclei. In reality, mitosis is a continuous process in which each event flows smoothly into the next. For discussion, however, it is convenient to look at mitosis — the M phase of the cell cycle — as a series of separate events: prophase, prometaphase, metaphase, anaphase and telophase.
The centrosomes determine the plane of cell division
Once the commitment to enter mitosis has been made, the cell enters S phase, and DNA is replicated. At the same time, in the cytoplasm, the centrosome(“central body”), an organelle that lies near the nucleus, doubles, forming a pair of centrosomes. In many organisms, each centrosome consists of a pair of centrioles, each one a hollow tube lined with nine microtubules. The two tubes are at right angles to each other. At the G2-to-M transition, the two centrosomes separate from each other, moving to opposite ends of the nuclear envelope. The orientation of the centrosomes determines the plane at which the cell will divide, and therefore the spatial relationship of the two new cells to the parent cell. This relationship may be of little consequence to single free-living cells such as yeasts, but it is important for cells that make up part of a body tissue. The material around the centrioles initiates the formation of microtubules which will orchestrate chromosomal movement. Plant cells lack centrosomes, but distinct microtubule organizing centers at either end of the cell serve the same role. The formation of microtubules leads to the formation of the spindle structure that is required for the orderly segregation of the chromosomes in cell division.
Chromatids become visible and the spindle forms during prophase
During interphase, only the nuclear envelope, the nucleoli, and a barely discernible tangle of chromatin are visible under the light microscope. The appearance of the nucleus changes as the cell enters prophase—the beginning of mitosis. Most of the cohesin that held the two products of DNA replication together since S phase has been destroyed, so the individual chromatids become visible. They are still held together by a small amount of cohesin at the centromere. Late in prophase, specialized three-layered structures called kinetochoresdevelop in the centromere region, one on each chromatid. These structures will be important in chromosome movements. Each of the two centrosomes serves as a mitotic center, or pole, toward which the chromosomes will move. Microtubules form between each pole and the chromosomes to make up a spindle which serves both as a structure to which the chromosomes will attach and as a framework keeping the two poles apart. The spindle is actually two half-spindles: Each polar microtubule runs from one pole to the middle of the spindle, where it overlaps with polar microtubules extending from the other half-spindle. The polar microtubules are initially unstable, constantly forming and falling apart, until they contact polar microtubules from the other half-spindle and become more stable. There are two types of microtubules in the spindle:
Polar microtubules have abundant tubulin around the centrioles. Tubulin subunits can aggregate to form long fibers that extend beyond the equatorial plate.
Kinetochore microtubules attach to the kinetochores on the chromosomes. The two sister chromatids in each chromosome pair become attached by their kinetochore to microtubules in opposite halves of the spindle. This ensures that one chromatid of the pair will eventually move to one pole and the other chromatid will move to the other pole. Movement of the chromatids is the central feature and accomplishment of mitosis.
Chromosome movements are highly organized
The next three phases of mitosis — prometaphase, metaphase, and anaphase — are the phases during which chromosomes actually move. During these phases, the centromeres holding the two chromatids together separate and the former sister chromatids move away from each other in opposite directions.
Prometaphase
Prometaphaseis marked by the disappearance of the nuclear envelope. The material of the envelope remains in the cytoplasm, however, to be reassembled when the daughter nuclei re-form. In prometaphase, the chromosomes begin to move toward the poles but this movement is counteracted by two factors:
Arepulsive force from the poles pushes the chromosomes toward the middle region or equatorial plate(metaphase plate) of the cell. The two chromatids are still held together at the centromere by cohesin. Thus, during prometaphase chromosomes appear to move aimlessly back and forth between the poles and the middle of the spindle. Gradually, the centromeres approach the equatorial plate.
Metaphase
The cell is said to be in metaphasewhen all the centromeres arrive at the equatorial plate. Metaphase is the best time to see the sizes and shapes of chromosomes because they are maximally condensed. The chromatids are now clearly connected to one pole or the other by microtubules. At the end of metaphase, all of the chromatid pairs separate simultaneously. This separation occurs because the cohesin holding the sister chromatids together is hydrolyzed by a specific protease, appropriately called separase. Until this point, separase has been present but inactive, because it has been bound to an inhibitory subunit called securin. Once all the chromatids are connected to the spindle, securin is hydrolyzed, allowing separase to catalyze cohesin breakdown. In this way, chromosome alignment is connected to chromatid separation. This process, called the spindle checkpoint, apparently senses whether there are any kinetochores that are unattached to the spindle. If there are, securin breakdown is blocked and the sister chromatids stay together.
Anaphase
Separation of the chromatids marks the beginning of anaphase, during which the two sister chromatids move to opposite ends of the spindle. Each chromatid contains one double-stranded DNA molecule and is now referred to as a daughter chromosome. What propels this highly organized mass migration is not clear. Two things seem to move the chromosomes along. First, at the kinetochores are proteins that act as “molecular motors.” These proteins, called cytoplasmic dynein, have the ability to hydrolyze ATP to ADP and phosphate, thus releasing energy to move the chromosomes along the microtubules toward the poles. These motor proteins account for about 75 percent of the force of motion. Second, the kinetochore microtubules shorten from the poles, drawing the chromosomes toward them. This shortening accounts for about 25 percent of the motion. During anaphase the poles of the spindle are pushed farther apart, doubling the distance between them. The distance between poles increases because the overlapping polar microtubules extending from opposite ends of the spindle contain motor proteins that cause them to slide past each other, in much the same way that microtubules slide in cilia and flagella. This polar separation further separates one set of daughter chromosomes from the other. The movements of chromosomes are slow, this slow speed may ensure that the chromosomes segregate accurately.
Nuclei re-form during telophase
When the chromosomes stop moving at the end of anaphase, the cell enters telophase. Two sets of chromosomes (formerly referred to as daughter chromosomes), containing identical DNAand carrying identical sets of hereditary instructions, are now at the opposite ends of the spindle, which begins to break down. The chromosomes begin to uncoil, continuing until they become the diffuse tangle of chromatin that is characteristic of interphase. The nuclear envelopes and nucleoli, which were disaggregated during prophase, coalesce and reform their respective structures. When these and other changes are complete, telophase — and mitosis — is at an end, and each of the daughter nuclei enters another interphase. Mitosis is beautifully precise. Its result is two nuclei that are identical to each other and to the parent nucleus in chromosomal makeup, and hence in genetic constitution. Next, the two nuclei must be isolated in separate cells which requires the division of the cytoplasm.
Cytokinesis: The Division of the Cytoplasm
Mitosis refers only to the division of the nucleus. The division of the cell’s cytoplasm, which follows mitosis, is accomplished by cytokinesis, which may actually begin before telophase ends. In different organisms, cytokinesis may be accomplished in different ways. The differences between the process in plants and in animals are substantial.
Animal cells usually divide by a furrowing of the plasma membrane, as if an invisible thread were tightening between the two poles. The invisible thread is actually microfilaments of actin and myosin located in a ring just beneath the plasma membrane. These two proteins interact to produce a contraction, just as they do in muscles, thus pinching the cell in two. These microfilaments assemble rapidly from actin monomers that are present in the interphase cytoskeleton. Their assembly appears to be under the control of Ca2+ released from storage sites in the center of the cell.
Plant cell cytoplasm divides differently because plants have cell walls. As the spindle breaks down after mitosis, membranous vesicles derived from the Golgi apparatus appear in the equatorial region roughly midway between the two daughter nuclei. Propelled along microtubules by the motor protein kinesin, these vesicles fuse to form new plasma membrane and contribute their contents to a cell plate, which is the beginning of a new cell wall. Following cytokinesis, both daughter cells contain all the components of a complete cell. Aprecise distribution of chromosomes is ensured by mitosis. Organelles such as ribosomes, mitochondria and chloroplasts need not be distributed equally between daughter cells as long as some of each are present in both cells; accordingly, there is no mechanism with a precision comparable to that of mitosis to provide for their equal allocation to daughter cells.
Дата добавления: 2016-07-18; просмотров: 2337;